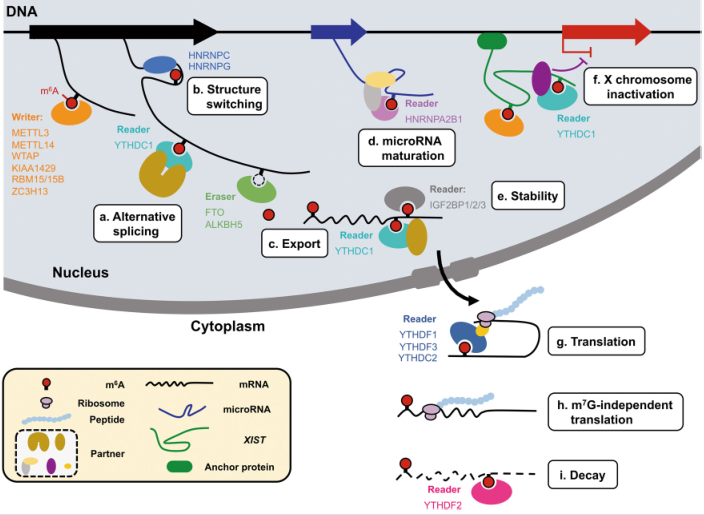
At present, there are 9 functions of reading proteins that regulate the m6A modification process. Today, we will not talk about it one by one, but mainly the YTHDF1 protein involved in the protein coding process. It mainly binds to the m6A site of mRNA in the brain. Neurodevelopment [1], dopamine secretion [2] and synapse formation [3] play an important role.

Article introduction:
On October 31, 2018, the Hechuan team of the University of Chicago, the Zhou Tao team of the University of Science and Technology of Shanghai and the Song Hongjun team of the University of Pennsylvania jointly published a paper in the Nature Journal, which first pointed out the effect of m6A modification on the learning and memory ability of mice. Today, Xiaobian will Explain this article in detail, and summarize the articles of Professor He Chuan on reading protein research in recent years.
Sample: wild type mouse, YTDHF1 knockout model mouse
Technical means: m6A RNA methylation sequencing , mRNA sequencing, CLIP sequencing and mouse cognitive-related functional experiments
Time point: initial stimulation period, LTP phase (late stimulation response) and secondary stimulation period
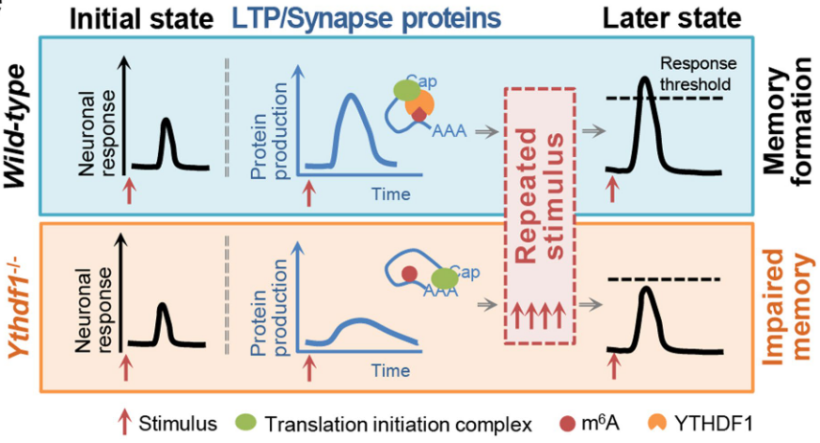
In this paper, the phenotypic response of the hippocampus in the YTHDF1 knockout and wild-type mice, the amount of related proteins and the changes of the anterior-posterior equipotential point threshold after LTP and secondary stimulation were studied.
At the initial stimulation, there was little difference in the phenotypic response of the two groups of mice; in the LTP phase, YTHDF1 knockout mice inhibited the synthesis of LTP-related proteins due to the lack of YTDHF1 protein, resulting in equipotential points when subjected to subsequent stimulation. Changes do not reach the threshold affects the ability to form memory.
1. YTDHF1 knockout mice are less sensitive to spatial memory and external stimuli
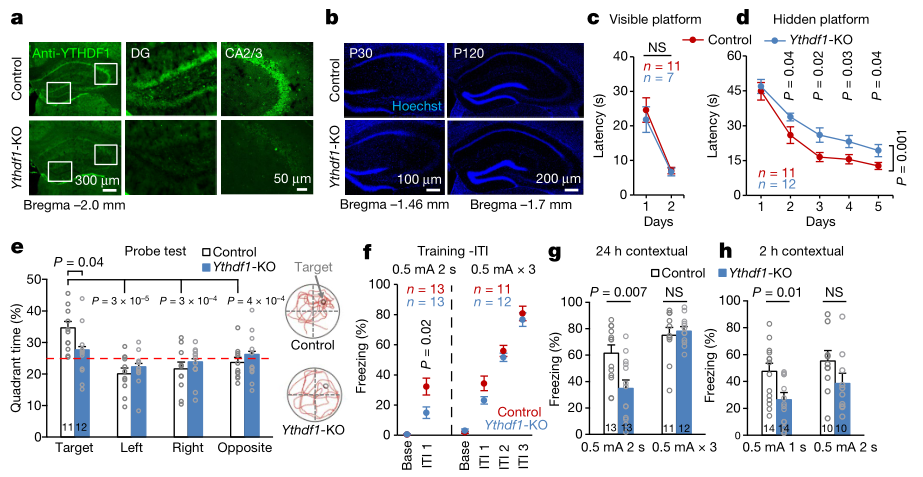
Previous experiments have confirmed that YTDHF1 is highly expressed in mouse hippocampus [4], so the authors constructed a YTDHF1 knockout model mouse (Fig. 1a) by the CRISPR-Cas9 technique and named it YTDHF1-KO mice. Compared with wild-type mice, the tissue morphology of the hippocampus in the KO group before the age of four years (Fig. 1b), the results of the transparent water maze training (Fig. 1c) were consistent with normal mice. Regardless of the water maze training (Fig. 1d), the probe experiment (Fig. 1e) and the sensitivity to external stimuli (Fig. 1f, g, h) are longer than the control, indicating the ability of knockout YTDHF1 protein for spatial memory and external stimuli. The response plays an important role.
2. YTDHF1 and related proteins participate in learning and memory of key LTP phases
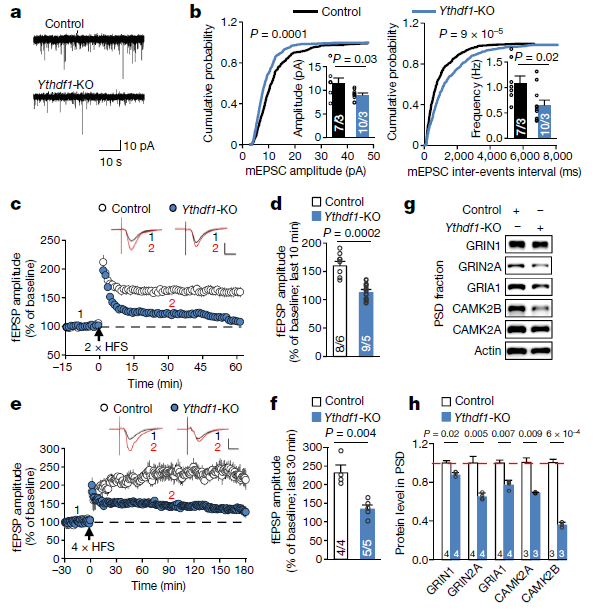
The authors then observed changes in the brain of the knockdown group of mice from physiological levels. The amplitude and frequency of synaptic currents in the knockdown group were lower than those in the control group (Fig. 2 a, b), and the morphology was smaller than that in the control group (Fig. 2 c, d). When exposed to external stimuli, the amount of change in the post-synaptic potential threshold of the nerve was lower than that of the control group (Fig. 2 e, f). At the same time, this process may be associated with five proteins associated with the post-synaptic density of the LTP phase (Fig. 2 g, h). The YTDHF1 complementation experiment showed that the above phenotype was restored.
3. YTHDF1 CLIP sequencing reveals that m6A peak is associated with synaptic transmission and LTP phase
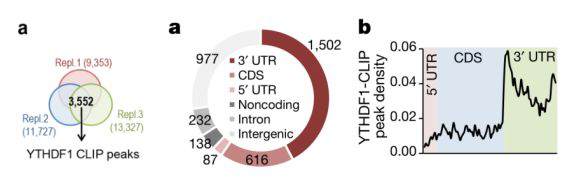
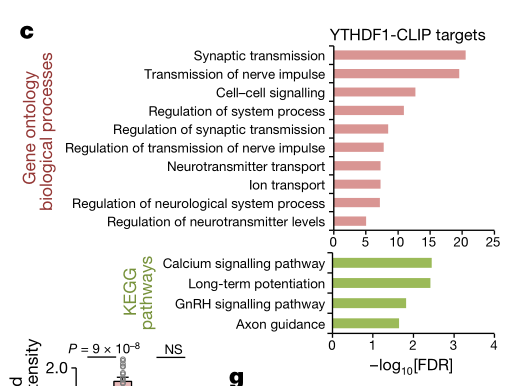
In order to further analyze the mechanism of action of YTHDF1 from a molecular perspective, the authors used YTHDF1-CLIP sequencing technology to detect all binding sites and m6A on YTHDF1 mRNA in mouse hippocampus (n=3, 4 mice mixed into one sample). Site. By taking the intersection of the m6A peaks of the three samples, a total of 3,552 peaks were found on 1,042 transcripts (Fig. 4a left). 1,502 peaks matched the 3' UTR region of the genome (Fig. 4a right); 67% of peaks matched the transcript and were enriched in the 3'UTR region (Fig. 4b). After commenting GO and KEGG on the above-mentioned peaks-related transcripts, it was found to be related to synaptic transmission and LTP phase, which coincided with the results of previous experiments.
4. m6A CLIP sequencing combined with YTHDF1 CLIP sequencing to screen for m6A sites that may bind to YTHDF1
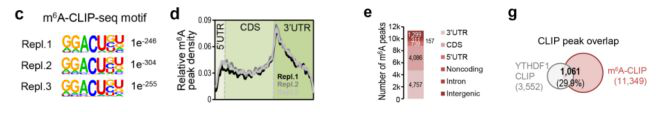
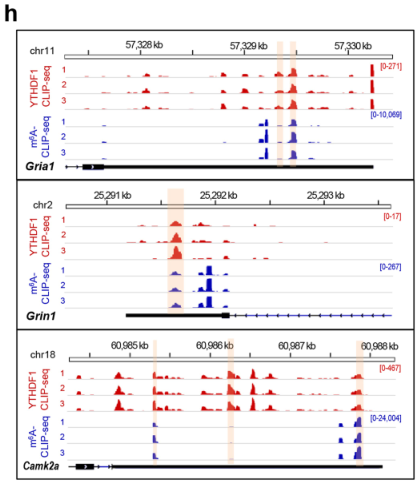
Next, the authors performed m6A CLIP sequencing on the same type of samples. After performing a motif analysis on each of the three samples, it was found that there were nearly 11,000 sequences in the three groups that were GGACU peaks (Fig. 5c). After comparing the peak to the transcript and the genome, it was found to be very similar to the YTHDF1 CLIP (Fig. 5d, e). After the intersection of the peaks of the two CLIP experiments, it was found that there were nearly 30% of the peak (Fig. 5 g), and the IGV results indicated that the m6A site of YTHDF1 and the m6A of synapse forming key proteins (such as Gira1, Grin1 and Camk2a, etc.) The sites are similar, and the two may bind to the m6A site to regulate protein synthesis (Fig. 5h).
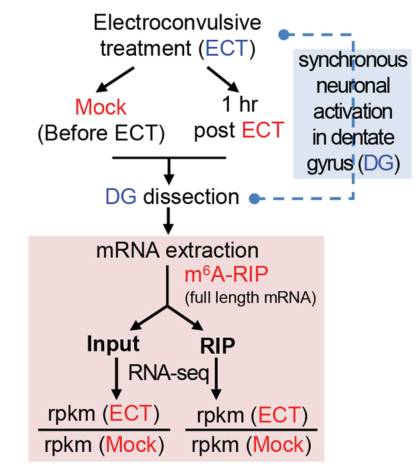
5. Firstly screened m6A RNA methylation profile in electroshock mice
Finally, the authors established an electroshock mouse model, which combined m6A RNA methylation sequencing and transcriptome sequencing on the dentate gyrus structure of the hippocampus, and quickly found many changes in the methylation expression of m6A. Genes for subsequent research.
to sum up:
This article demonstrates that m6A RNA methylation in mouse hippocampus regulates learning and memory production and can influence the regulation of key target genes involved in neuronal formation, providing further insight into m6A modification in learning and memory analysis mechanisms. I know.
Professor He Chuan's m6A RNA methylation Reader related protein results summary:
1. 2013.11 Nature IF: 41.58
Binding proteins with a YTH domain affect function through mRNA binding.

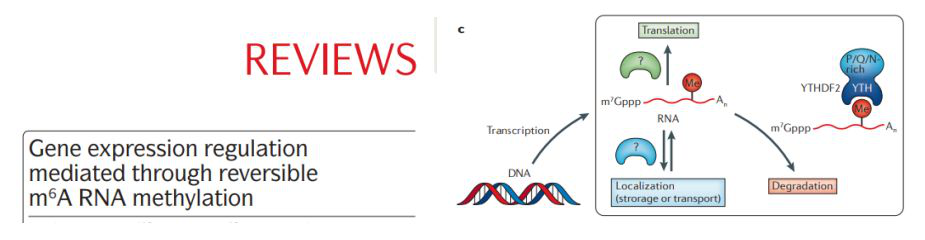
2. 2014.5 Nature IF: 41.58
Explain the mechanism of action of the Reader protein.
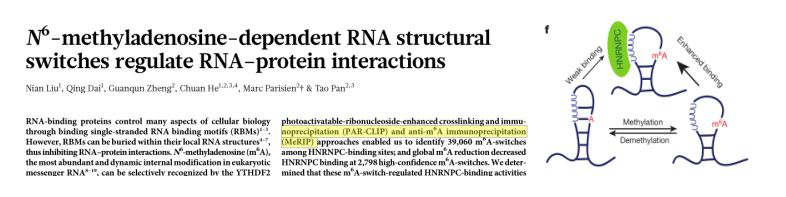
3. 2015.2 Nautre IF: 41.58
The biological function of m6A methylation binding protein HNRNPC binding to LncRNA is revealed.

4. 2017.4 Cancer Cell IF: 22.84
The biological function of m6A methylation binding protein HUR binding to LncRNA is revealed.

references:
1. Hess, ME et al. The fat mass and obesity associated gene (Fto) regulates activity of the dopaminergic midbrain circuitry. Nat. Neurosci. 16, 1042-1048 (2013).
2. Weng, Y.-L. et al. Epitranscriptomic m 6 A regulation of axon regeneration in the adult mammalian nervous system. Neuron 97, 313-325.e6 (2018).
3. Li, L. et al. Fat mass and obesity-associated (FTO) protein regulates adult neurogenesis. Hum. Mol. Genet. 26, 2398-2411 (2017).
4. Lein, ES et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168-176 (2007).
Yunxu Bio provides exclusive one-stop service for RNA methylation sequencing in China
Cloud-sequence organisms provide colorimetric detection of overall m6A methylation modification levels, RNA methylation sequencing, MeRIP-PCR validation, and RIP, RNA Pull Down mechanism research services. RNA methylation sequencing technology is the real implementation of m6A, m5C and m1A modification, detection of molecules in addition to mRNA, but also detection of circular RNA, LncRNA and other non-coding RNA. Since 2016, the sample size has accumulated more than 1000+, and the MeRIP rich integration power is over 98%.
Now, in order to solve the problem of small sample size of customers, ultra-micro-MeRIP sequencing technology is introduced, and 500 ng of total RNA can be used for sequencing experiments. Special samples can also be used for RNA methylation sequencing, such as serum, plasma, exosomes, and paraffin samples.
Cloud sequence biological RNA methylation product list:
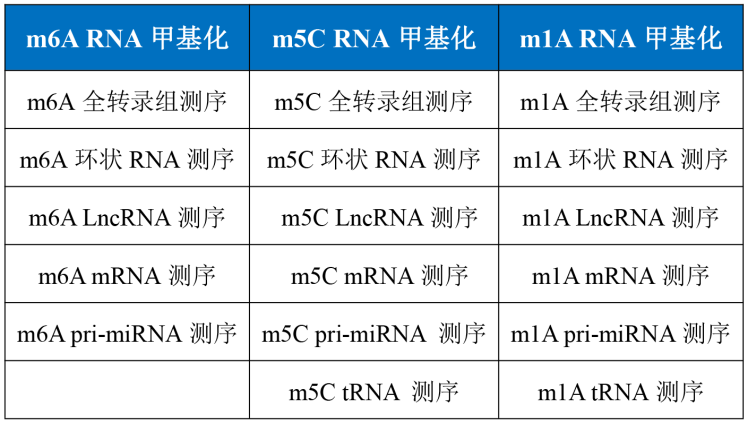
Product Links:
m6A RNA whole transcriptome methylation sequencing
m5C RNA whole transcriptome methylation sequencing
M1A RNA transcriptome methylation sequencing colorimetric assay for overall methylation level ultra-micro-MeRIP sequencing
RIP
RNA Pull Down
Shanghai Yunxu Biological Technology Co., Ltd.
Shanghai Cloud-seq Biotech Co., Ltd.
Address: 3rd Floor, Building 20, No. 518, Zhangzhu Road, Songjiang District, Shanghai 
Telephone Fax Website:
mailbox:
Fast Shipping Laser Distance Sensor
Hi there,
We create a new payment method online for you, so that all of our customers, and you can get the samples quickly. With the competitive prices online, it is easy to operate and finish your sample orders of JRT Laser Distance Sensor. It's really amazing, right? Let's go!
Greetings

sensor lidar,laser measurement,laser measuring equipment,range of radar
Chengdu JRT Meter Technology Co., Ltd , http://www.accuracysensor.com